Frameworks to assess the acceleration of biomedical research using GPAI
GPAI capability framework
To analyze the potential for GPAI-driven acceleration of biomedical research, we first developed a framework of GPAI capability levels. Drawing on multiple existing frameworks35,36,37,38 and analyzing biomedical research requirements, we synthesized a simple, unified framework of GPAI research capability with two key dimensions:
-
Cognitive capability, encompassing research activities primarily involving information processing, analysis, and decision-making, including literature review, hypothesis generation, experimental design, data analysis, result interpretation, and manuscript preparation.
-
Physical capability, involving laboratory procedures, experimental setup, and material handling, which are made possible by robotics, lab automation, and automated experiment execution.
For both cognitive and physical capabilities, we define three levels that chart the progression of GPAI integration into the research process:
-
At the “No GPAI” level, humans perform all work manually, possibly assisted by non-GPAI tools.
-
“Next-level” capabilities represent GPAI systems that partially automate the research cycle but require significant human intervention, as demonstrated by several current systems.
-
“Maximum-level” capabilities represent a radically transformed future scenario with advanced capabilities and high-level autonomy where GPAI conducts expert-level research with minimal to no human supervision.
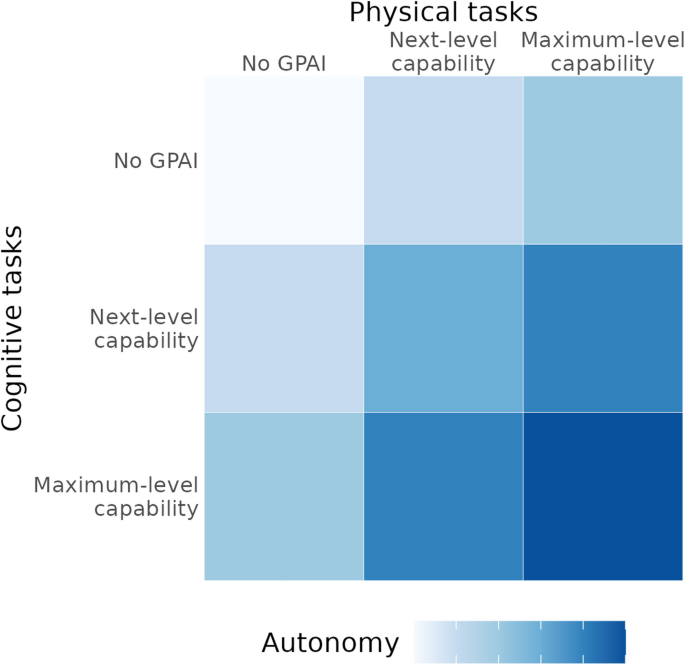
The joint development of cognitive and physical capabilities lead to increases in the emergent attribute of autonomy, i.e., the ability to operate independently across the complete research cycle. High levels of autonomy are key to the most radical acceleration of research (Fig. 1).
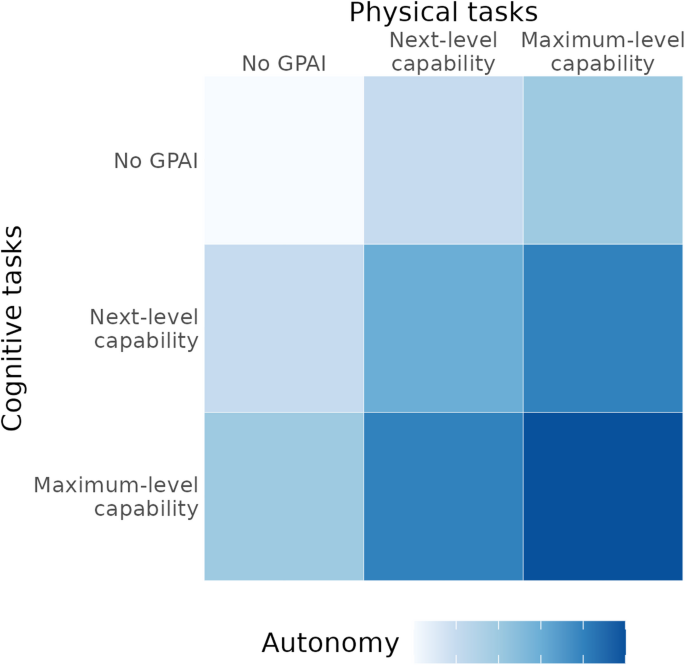
Adapted from Tom et al.38.
Capability framework illustrating how cognitive and physical capabilities of GPAI combine to yield autonomy. Higher autonomy levels enable increasingly significant acceleration of the biomedical research process.
Research task framework
Drawing on established concepts from the literature39,40,41,42,43, we developed a structured, end-to-end framework of the biomedical research process comprising nine major research tasks (Fig. 2), as well as constituent sub-tasks (Table S1). This allows us to map specific GPAI capabilities found in the literature to individual research tasks, track the varying levels of automation possible across different aspects of research, and better identify bottlenecks and opportunities for acceleration.
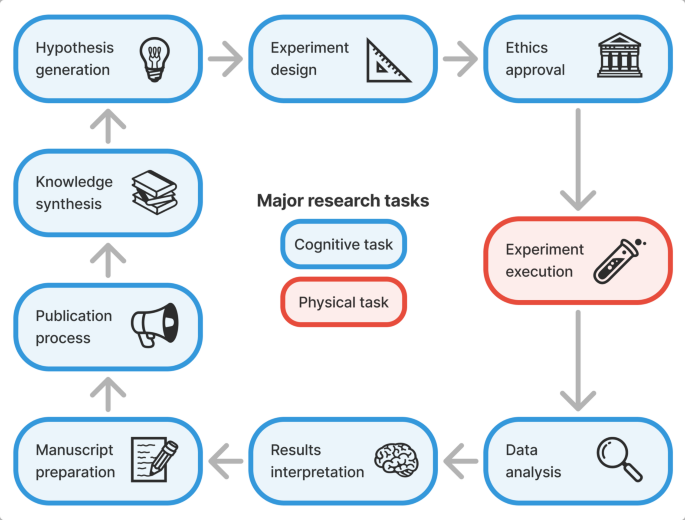
End-to-end biomedical research task framework covering nine major tasks. Blue boxes indicate cognitive tasks, the red box the sole physical task (experiment execution), and arrows depict the typical order and iterative nature of research projects.
The major research tasks in our framework consist of the following:
-
1.
Knowledge synthesis encompasses the collection, critical evaluation and integration of scientific information. Recent GPAI approaches achieve human-level or higher precision in literature-based tasks and operate at high throughput29. Systems automate literature search, evaluation, and summarization21,44 while demonstrating critical thinking and evaluation capabilities27,45.
-
2.
Idea & hypothesis generation involves identifying promising research problems, formulating testable hypotheses, developing theoretical frameworks, and assessing feasibility. GPAI systems are increasingly able to develop new research ideas by analyzing patterns in existing literature and proposing candidate hypotheses and mechanisms28,33,46. Advanced systems can autonomously generate and, in some cases, rank candidate hypotheses by novelty and feasibility, then refine key research questions into actionable, experimentally testable goals26,27,44,47.
-
3.
Experiment design includes method selection, protocol development, and planning data collection, including quality control. GPAI systems now demonstrate capabilities in suggesting experimental methods and designing wet-lab or computational protocols27,28,48, with advanced systems able to formulate experimental plans, predict and optimize parameters, and integrate quality control and reproducibility measures, including protocol standardization21,24,44,49,50.
-
4.
Ethical approval & permits involve initial screening of research proposals, scientific review, ethics assessment, regulatory compliance, and administrative processing. While currently the most human-centered process, GPAI is beginning to assist in areas such as automated screening for ethical issues, documentation preparation, and compliance checks51,52,53. GPAI systems have potential to enhance informed consent processes54,55, improve scientific review efficiency, and support compliance checks against applicable regulations and guidelines56.
-
5.
Experiment execution represents the physical dimension of research, including preparation, execution, documentation, troubleshooting, material management, and equipment maintenance. Self-driving laboratories can execute fully automated multi-step workflows with parallel samples, achieving significant acceleration through continuous operation, exception handling and adaptive optimization24,30,57,58,59. Advanced robotic systems execute protocols, monitor experimental conditions, and manage samples while improving throughput and reducing time requirements24,31,60,61.
-
6.
Data analysis encompasses collection, cleaning, statistical analysis, visualization, and validation. GPAI systems increasingly automate much of the analysis pipeline, from preprocessing to statistical modeling and interactive exploratory analysis49,62,63. Modern systems collect and integrate diverse data types and automate data cleaning and preprocessing, with high agreement with expert analyses within end-to-end automated workflows21,27,44,64.
-
7.
Results interpretation involves synthesizing experimental findings, evaluating hypotheses against results, and contextualizing findings within broader scientific knowledge. GPAI systems can now integrate computational and experimental data to connect predictions with outcomes and employ multi-agent approaches to critically evaluate results21,27,28. Advanced systems can compare experimental results to hypotheses and contextualize findings within existing knowledge29,44.
-
8.
Manuscript preparation includes documenting methods, presenting results, managing references, creating figures and tables, facilitating data sharing, and handling writing and revision. GPAI systems can now generate complete manuscript drafts with methodological descriptions and iteratively improve them through automated feedback45,65. Modern systems document experimental methods, manage references, generate figures, and prepare shareable data and code26,29,44,49,66.
-
9.
Publication process involves journal selection, submission, screening, peer review, and revision. GPAI assistance is emerging in areas such as journal and reviewer recommendation, formatting for submission, and generating peer reviews67,68,69,70. GPAI-generated feedback approximates human reviewers, and editorial experiments have found AI reviews sufficiently accurate to be helpful; GPAI can identify several (though not all) mathematical and conceptual errors71,72,73.
Each of these main tasks comprises several subtasks with varying potential for GPAI acceleration. While these main tasks usually proceed in the order presented, research projects often require iterative transitions between tasks, such as when feedback from reviewers requires a return to experimental design or when unexpected results necessitate revisiting hypothesis generation. Notably, experiment execution represents the physical dimension of research, while all other tasks are primarily cognitive. This framework forms the basis for our investigation of how different levels of GPAI capabilities can accelerate the biomedical research process.
Integration of GPAI capability and research task frameworks
Combining the frameworks introduced above yields a matrix of research acceleration scenarios across GPAI capabilities and major research tasks (Table S2).
This integrated framework allows us to systematically analyze how different levels of GPAI capability transform the research process, identify where the greatest potential for acceleration exists, and what bottlenecks might remain even with advanced GPAI systems.
Evidence for acceleration potential
To obtain concrete GPAI acceleration factors, we conducted a scoping review of literature on GPAI accelerating research. It reveals significant variation in potential speedups between cognitive and physical tasks, with cognitive tasks generally showing higher acceleration factors21,24,29,60. The evidence ranges from modest improvements to dramatic transformations in research timelines.
Acceleration of cognitive tasks
The strongest empirical evidence for cognitive task acceleration comes from recent applications of GPAI in research environments, though the measurement approaches and reported metrics vary across studies. For instance, recent industry reports estimate GPAI-driven research and development in drug discovery from initial research to the preclinical stage, yielding a 1.3-2x efficiency increase, corresponding to a 25–50% reduction in cost and time74. Analysis of flow cytometry data in clinical immunology has been accelerated 2-4x by GPAI, reducing the required time from 10 to 20 min to 5 min, while maintaining expert-level accuracy62.
Looking at related tasks outside the core of science, studies have reported a 1.3x speedup in consulting tasks75, a 1.7x speedup (40% time decrease) in professional writing tasks76, and a 2.2x speedup (55% time decrease) for coding tasks, according to GitHub’s internal report on Copilot77.
The reported acceleration potential increases significantly with more advanced GPAI systems and setups optimized for automation. For example, in scientific knowledge synthesis, PaperQA2 demonstrated an ~ 75-300x speedup, writing high-quality Wikipedia-style articles in 8 minutes29 (a task that human editors report taking 10–40 hours78. In bioinformatics, a GPAI agent capable of fully automated multi-omic analyses reportedly just required 5 min for the exemplary task of identifying differentially expressed genes between bulk RNA-seq samples49. Compared to the 10–12 hours79,80 reported by two bioinformatics facilities for the same type of task, this represents an approximate speedup of ~ 120-140x.
Extending to complete research cycles, a GPAI agent team claimed to have developed SARS-CoV-2 nanobodies in a fraction of the time human researchers would have needed27,81. Another automated biomedical GPAI system called BioResearcher reported achieving a ~ 150-300x speedup by completing full dry lab research cycles, from literature searches to the execution of computational experiments, in approximately 8 h versus the traditional 7–14 weeks21. Similarly, Sakana AI’s ‘AI Scientist’ demonstrated high efficiency through full automation in computer science research, capable of exploring research ideas in ~ 15 min (a rate of ~ 50 ideas in 12 h)26.
Acceleration of physical tasks
In laboratory settings, physical task acceleration also shows promising results, though generally with somewhat lower acceleration factors than purely cognitive tasks. An integrated robotic chemistry system achieved a 1.7x speedup in synthesizing nerve-targeting agents compared to manual methods, completing the entire 20-compound library in 72 h instead of 120 h, reportedly with comparable quality82.
Protein engineering with integrated GPAI testing and feedback demonstrated a reduction in project duration from 6 to 12 to six months (1-2x speedup) in real-life testing (including shipping delays). The authors suggest that with better planning it could be reduced to two months (3-6x speedup) and in the best-case scenario of continuous operation to just 1–2 weeks (~ 15-50x speedup)24. A GPAI-driven microbial culturomics platform overcomes the variability of manual methods by using imaging to autonomously inform colony selection, yielding a more than 20x speedup by achieving an isolation throughput of 2,000 colonies per hour in an integrated pipeline60. An automated materials discovery platform integrated ML screening, robotic synthesis, and characterization, reportedly reduced material sintering times from 2 to 6 h to 10 min (12-36x speedup) and was noted to reduce entire processes from hours or days to minutes83.
An automated chemical workflow handling 16 parallel samples conducted 688 experiments in 8 days. Compared to manual methods, which were estimated to take half a day per experiment, this represents a ~ 40x speedup. In addition to these results, the study reported estimated acceleration factors of ~ 10x–100x compared to conventional workflows, where the lower range corresponds to semi-automated methods and the higher end to manual approaches30. Self-driving laboratories that integrate robotics, additive manufacturing, and GPAI were projected to accelerate materials and molecular discovery by 10–100x through combining gains from robotics (2x), active learning (5-20x), process intensification (up to 100x) and continuous operation (2–3 ×)84.
Based on internal industry data, one prominent cloud lab suggests its GPAI implementation enables a 2x speedup in time-to-publication (from an average of 1.96 years to one year) and claims to generate publication-quality data 90x faster by reducing traditional 3-month timelines to 24 hours85. In a notable anecdote, a PhD student reported replicating years of their previous project’s work in just one week using an automated robotics platform with 24/7 operation (~ 100x speedup)86.
Challenges and constraints of biomedical research acceleration
The empirical evidence reviewed above highlights the potential for GPAI to accelerate both cognitive and physical research tasks, with some studies demonstrating order-of-magnitude improvements. These impressive figures often reflect optimized scenarios or specific sub-tasks. Realizing such acceleration consistently across the entire research lifecycle is subject to various practical, biological, infrastructural, and institutional constraints.
One major category of reported constraints relates to the implementation and operation of automated systems and self-driving laboratories. Studies note that creating robust self-driving laboratories requires significant investment and complex integration of automated experiments with GPAI decision-making87,88. Even once operational, researchers report challenges in automated or cloud lab environments including remote troubleshooting difficulties, reduced experimental flexibility for exploratory research, and limitations in applicability for academic settings characterized by frequent directional pivots89. Comparing the effectiveness of different self-driving laboratories is also reportedly difficult due to challenges in defining standardized performance metrics that capture the nuances of diverse lab setups88. Finally, translating discoveries made in controlled self-driving laboratory environments to real-world applications faces hurdles related to storage stability, limited resources, and self-sufficiency without expert supervision90.
Another set of limitations highlighted in the literature concerns data and the GPAI models themselves. The performance of models central to GPAI and self-driving laboratories can be hampered by shortcomings in available data, such as the common lack of negative results or detailed metadata in published literature91. Building robust GPAI decision-making models often requires large, high-quality, information-rich datasets, the generation of which can be a bottleneck92.
Perhaps the most fundamental constraints reported are inherent biological and physical limits. While automation can speed up workflows like liquid handling or data acquisition, the underlying biological processes often have irreducible timescales89. Cell-based experiments, for instance, remain resource-intensive and subject to variability, with limits imposed by factors like maximum cell growth rates under specific conditions92. Self-driving laboratories optimize experiments around these biological components rather than altering their intrinsic limits87,91. Even the speeds of biochemical processes, like enzyme kinetics or protein folding, impose natural limits that automation cannot bypass90. Finally the complexity of biology also presents a challenge, as fully understanding and predicting cellular behavior remains difficult without comprehensive perturbation data, even when advanced models are utilized89,92.
Aside from the technical and biological hurdles, significant limitations arise from established social and institutional structures and processes. Although GPAI can accelerate certain tasks, the overall pace can be slowed by procedural delays inherent in the current academic system, as they may not scale readily with technological advancements. Ethics approval processes average between 50 and 138 days93. Similarly, publication faces substantial delays: preprint-to-publication averages 199 days94, with submission-to-publication times within journals ranging from 91 to 639 days95. The peer review process itself was reported to take 17 weeks in one study96, despite reviewers spending only about six hours per review in each round97.
These documented limitations, which have been identified in the literature alongside the potential for acceleration, suggest that achieving maximum theoretical acceleration across entire research workflows poses significant practical, technological, and ethical challenges. In fact, even beyond these operational and inherent limitations, rigorously assessing the extent of GPAI-driven acceleration itself presents methodological complexities. The interpretation of reported accelerations requires clear baseline values and system boundaries. Highly task-specific accelerations, such as GPAI agents that rapidly design nanobodies27 or automate the planning and coding of bioinformatics analyses21,49, are valuable but must be distinguished from reductions in overall project duration.
Assessing the acceleration of research driven by GPAI requires methodological rigor, as the perceived benefits depend on the chosen benchmark (e.g., humans, optimized laboratories, or state-of-the-art automation). For example, robotic systems accelerate workflows such as chemical synthesis30 or high-throughput screening86 primarily through parallelism and continuous operation, rather than pure task speed. Such throughput gains are significant, but must be compared with appropriate advanced baselines—not just sequential execution by humans—and must take into account fixed setup costs that can reduce the benefit of switching to automated workflows, especially in isolated or small-scale projects.
Similarly, the acceleration offered by GPAI in the computer-aided discovery of therapeutics33 or proteins24 can only be judged when the validation effort for GPAI-generated hypotheses is taken into account. Therefore, precise reporting standards are crucial, requiring transparency regarding benchmarks, system boundaries, the distinction between actual speed and throughput, and full consideration of all operating and validation costs in order to accurately assess the benefit of GPAI.
Acceleration factors with GPAI levels
Our literature review reveals a bimodal distribution of acceleration factors across research tasks, with most observed values clustering either at lower levels (below 3x) or higher levels (above 10x for physical tasks and above 50x for cognitive tasks) (Fig. 3, Table S3). This pattern suggests two distinct regimes of reported GPAI-driven acceleration: incremental acceleration attainable today across the entire research process (Next-level GPAI), contrasted with transformative acceleration, achievable only through advanced systems and configurations (Maximum-level GPAI), and currently restricted to specific research tasks—thereby directing attention toward the theoretical maximum of research acceleration.
To translate these empirical findings into practical modeling scenarios, we assigned two distinct acceleration profiles to the capability levels defined in our framework:
-
1.
Next-level GPAI: This profile models the current acceleration potential of current GPAI systems as they diffuse through the research ecosystem. Based on the lower cluster of our empirical findings, we estimate acceleration factors of 2× for both cognitive and physical tasks—a mid-range value from observed current improvements (Fig. 3) We assume that these represent realistic, immediately achievable improvements that organizations can expect when implementing current GPAI and lab automation technologies, as already demonstrated in preclinical drug discovery for pulmonary fibrosis33.
-
2.
Maximum-level GPAI: This profile explores the transformative acceleration potential of research with future, highly advanced GPAI systems. Drawing from the upper cluster of documented capabilities, we estimate acceleration factors of 100× for cognitive tasks and 25× for physical tasks (Fig. 3). We deliberately selected to err on the conservative side by picking acceleration factors from the lower end of reported values for physical tasks (where more empirical evidence exists) and below even the minimum observed value for cognitive tasks (where evidence is more limited). While these factors may seem extraordinary, they represent acceleration potentials that have been demonstrated in specific contexts and thus provide evidence for possible future scenarios.
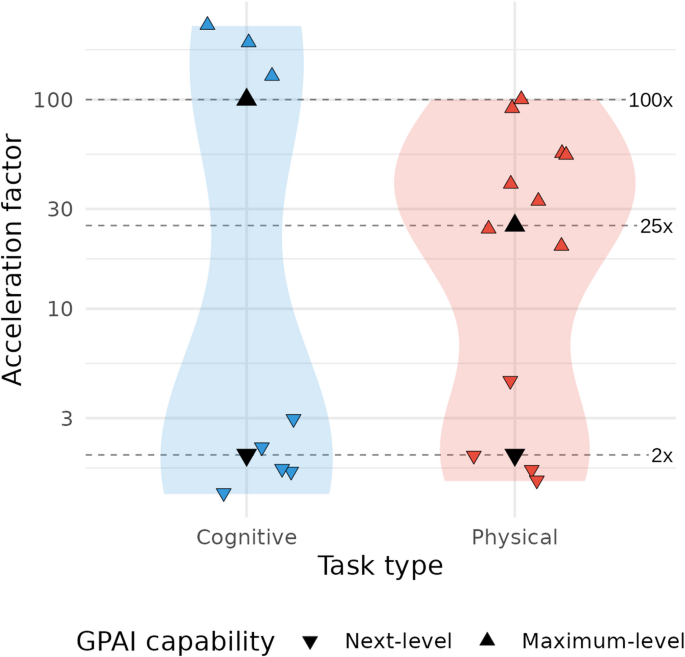
Reported acceleration factors for cognitive and physical research tasks. Violin plots depict the distribution of 20 acceleration factors on a log10-scale extracted from 16 publications for cognitive (blue) and physical (red) tasks. Individual studies are shown as jittered triangles; downward symbols (▼) indicate next-level capability, upward symbols (▲) indicate maximum-level capability. Four black triangles and three dashed lines (at 2×, 25×, 100×) denote the factors chosen for our modeling scenarios.
Biological time constants in acceleration modeling
It is important to note that the acceleration factors cited above derive primarily from high-throughput in-vitro experiments and computational tasks. However, biomedical research, particularly involving whole organisms, contains certain irreducible processes that cannot be accelerated beyond natural biological limits (such as time for cell growth, animal model development, or tumor progression). We therefore include a “non-compressible” time constant in our model representing irreducible intervals dictated by biological processes that remain fixed regardless of technological advancement.
Acceleration scenario modeling
Taken together, we propose the following simple formula to estimate research time:
Total research time = (Compressible time ÷ Acceleration factor) + Non-compressible time.
To demonstrate the effect of different acceleration scenarios on research time, we model a hypothetical 3-year biomedical research project representative of a typical PhD project duration. In this example we assume 24 months of cognitive work and 12 months of physical experimental work, of which 3 months represent biological time constants that cannot be compressed (Table 1).
Cognitive tasks offer greater potential for GPAI-driven acceleration due to their longer initial durations and higher acceleration factors, lacking the inherent limitations of biological processes and requiring less infrastructural investment. Still, achieving the most significant acceleration across biomedical research depends on maximizing GPAI capabilities in both cognitive and physical domains.
Applying maximum-level GPAI to both cognitive and physical tasks in our example reduces a 3-year (36-month) biomedical research project to 3.6 months, a 10x acceleration despite the biological time constant. While this constant constitutes a small fraction of the total timeline without GPAI (3 of 36 months ~ 8.3%), it becomes the dominant factor (3 of 3.6 months ~ 83%) under maximum-level acceleration.
Fields heavily dependent on in-vitro or computational approaches may realize acceleration factors approaching our maximum estimates, while those requiring extensive in-vivo work will experience more modest overall timeline reductions due to the presence of larger biological time constants.
Exploratory expert elicitation
In order to assess both the plausibility of our acceleration factors and limiting conditions, as well as the prevailing attitude and expectations of biomedical researchers toward GPAI for accelerating research, we conducted an exploratory elicitation with eight biomedical expert researchers. They (1) estimated time allocation across nine research tasks, (2) evaluated the plausibility of maximum-level acceleration factors (∼100× cognitive, ∼25× physical) for each task on five-point scales, (3) rated potential limiting factors on research acceleration, and (4) provided open-ended considerations.
The experts reflected on how our findings would apply to projects they had led from conception to publication in high-impact journals. They reported an average project duration of 72 months (Fig. 4), which is twice as long as our hypothetical example, but is consistent in the proportional distribution between cognitive and physical tasks: cognitive tasks took up 73% of project time (52 months), which is similar to the 67% in our hypothetical project. Experts identified “experiment execution,” “publication process,” and “data analysis” as the most time-consuming research tasks, while “ethics approvals and permits” and “knowledge synthesis” were rated as the least time-consuming.
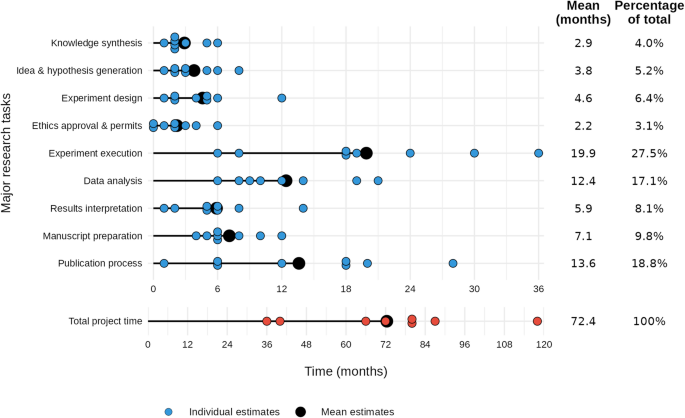
Timeline of major research task durations across studies. Horizontal dots show individual estimates (one dot per study) of time spent (in months, on a 6-month–interval axis) on each major research task, with lollipop-plots indicating the task’s mean. On the right, for each task, the rounded mean duration (months) and percentage of the total project time is shown. Below, a separate row presents individual and mean estimates for overall project duration (in months, on a 12‐month–interval axis).
When evaluating our estimates for maximum-level acceleration (~ 100-fold cognitive, ~ 25-fold physical), biomedical experts judged experiment design and execution, and hypothesis generation to be strongly overestimated, while greater acceleration potential was deemed plausible for administrative tasks. Respondents consistently considered our acceleration estimates overestimated for experimental design (7/7 responses), experimental execution (7/8) and hypothesis generation (7/8). In contrast, experts considered high acceleration factors plausible for structured administrative processes: ethics approval (4/7), manuscript preparation (4/8), and publication processes (3/8), with the rest of the responses mixed between over- and underestimation (Fig. 5).
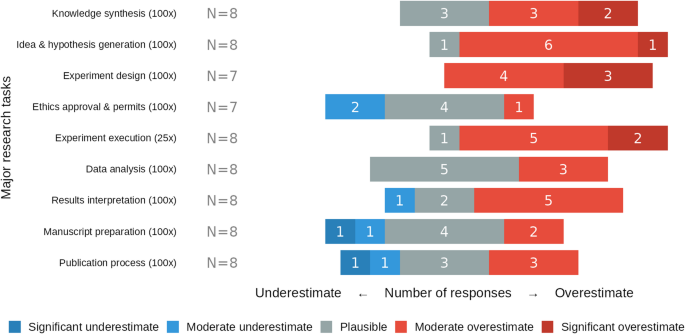
Perceived plausibility of maximum-level GPAI-acceleration factors across the nine major research tasks. Colors denote responses: significant underestimate (dark blue), moderate underestimate (light blue), plausible (grey), moderate overestimate (light red), and significant overestimate (dark red), with number of responses in white.
We asked experts to rate the significance of various potential bottlenecks (Fig. 6). While many factors showed a mixed response, scientific community assimilation was rated by all respondents as a moderate (2/8 responses), major (4/8) or crucial limit (2/8). In contrast, human strategic direction was seen as a lesser constraint, with a majority rating it as a minor (4/8) or insignificant limit. There was a marked consensus that stakeholder coordination is only a moderate limit (5/8).
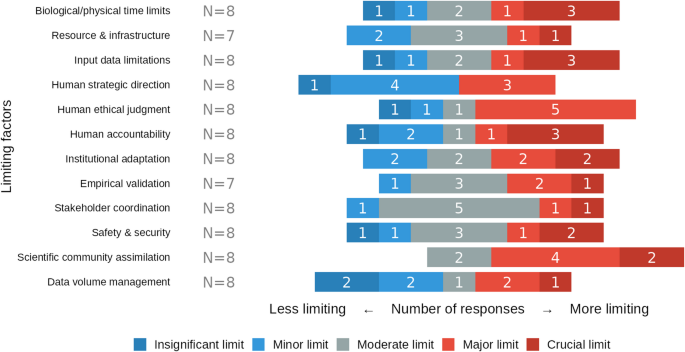
Perceived severity of factors that may limit GPAI-driven research acceleration. Colors encode response categories: insignificant limit (dark blue), minor limit (light blue), moderate limit (grey), major limit (light red), and crucial limit (dark red), with number of responses in white.
In addition to the quantitative ratings, experts provided general considerations (Table S7), in which they highlighted irreducible biological and social constraints. One researcher noted that for their project, “the blood sampling of 200 individuals simply takes a definite time,” while another pointed to the “fundamental time-frame of the experiment (i.e. looking at 3 month effect after intervention).” The limitations of social processes were also stressed, “the speed of publication with peer review and also the response of the co-authors cannot be changed,” underscoring that institutional adaptation and human coordination remain important bottlenecks. Experts also emphasized practical challenges in system integration and the socio-economic barriers to adoption. One expert noted the difficulty of “interfacing of various output/input systems” and also pointed to the “Cost/Benefit ratio,” suggesting that the high upfront cost requires “phenomenal trust in results” (see Tables S4–S7 for full survey results and Information S8 for survey interface).
link






More Stories
McConnell secures $2.6B in federal funding for Kentucky, boosting Louisville redevelopment and research
North America Pre-clinical Scientific Research Medical Device Market Report 2033
75 years ago, her cells were taken without consent. Today her family honors her legacy to make sure it never happens again